By: Elaina Graham
Hi! I am finishing my second year as a PhD student at USC in Dr. John Heidelberg’s lab. I am a microbial ecologist who looks at genes directly from the environment to explore the diversity of microbial life, and I’m also a WIES Fellow at Catalina this summer.
Microbes truly rule the world (or at least the ocean!) because they help mediate all of the major nutrient cycles in the ocean by helping to turnover carbon, nitrogen, and even sulfur. In previous research, we’ve found a very interesting organism that falls into a group called ‘Aerobic Anoxygenic Phototrophs’ (AAnP) – it uses light to produce energy, but has never before been shown to convert carbon sources like CO2 into organic carbon sources that can be used by the cell. This organism was seen all across the global ocean, in samples we looked at from the North Pacific, North Atlantic, and Mediterranean Sea. Our goal is to determine if it is now present in the waters around Catalina Island.
Because around 99% of the microbes in the ocean haven’t been cultured yet, it is important for us to find other methods to look at the diversity of microbes in the ocean.

Aboard the RV Yellowfin, getting ready to collect water. Each bottle on the rosette you see here can collect 12 Liters of water
Spending the summer out at Wrigley has been an amazing experience thus far because it has given me direct access to seawater from the cove that I can use to look for signs of the organism. Because these particular bacteria can make up anywhere from 1% -11% of the community, we often need a lot of water to look for the genes indicating the presence of this group. I have largely been sampling in the cove, but also had a wonderful opportunity earlier in the summer to go on USC’s boat RV Yellowfin (with a class I was teaching out at Wrigley prior to starting my summer research). This helped really kick off my project by enabling me to not just collect surface waters in the cove, but also sample by vessel for water that is 100 meters below the surface!
Once I got this water, I filtered it and began using something called PCR to start looking for signs of AAnP microbes. The method of PCR searches the environmental DNA for a specific gene, and once it finds this gene it amplifies it and produces billions of more copies – enough that we can ultimately sequence. Doing this requires you to be extra clean because even the tiniest bit of contamination from dust in the air or a cough could ruin the experiment. To avoid this we do these experiments in a special clean hood.

(Left) The PCR clean hood we use to do our molecular work and keep contamination out. (Right) The machine we use to run PCR by cycling through hot cycles up to 96°C.
While doing this I found myself getting very low amounts of ‘positive’ results, which could either have been because A) our group was missing from Catalina waters or B) there were issues with the PCR. Because I had seen previous studies at the San Pedro Ocean Time Series (SPOT) indicating the presence of these organisms, I decided I needed some kind of positive control. So I recreated a classical experiment where multiple AAnP bacteria were isolated from algae – I chose to use Sargassum horneri. AAnP bacteria tend to be pink, orange, or yellow due to the pigments they use to acquire light, so if they grow on solid media they are quickly identified!
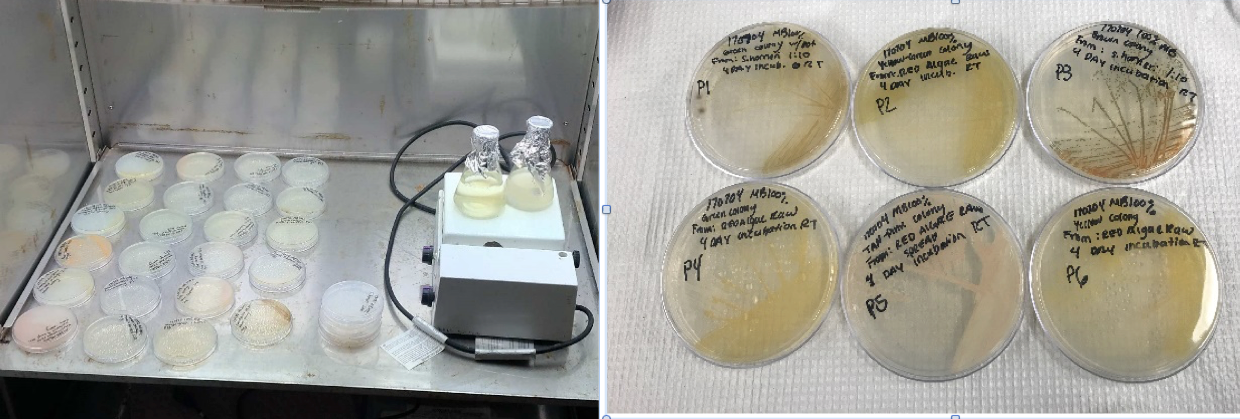
(Left) My incubator setup for culturing bacteria. I have solid agar plates where I isolated pigmented bacteria from the algae, as well as liquid media where I am using algae to grow up a culture of its associated bacteria. (Right) Close up of some of these cultures
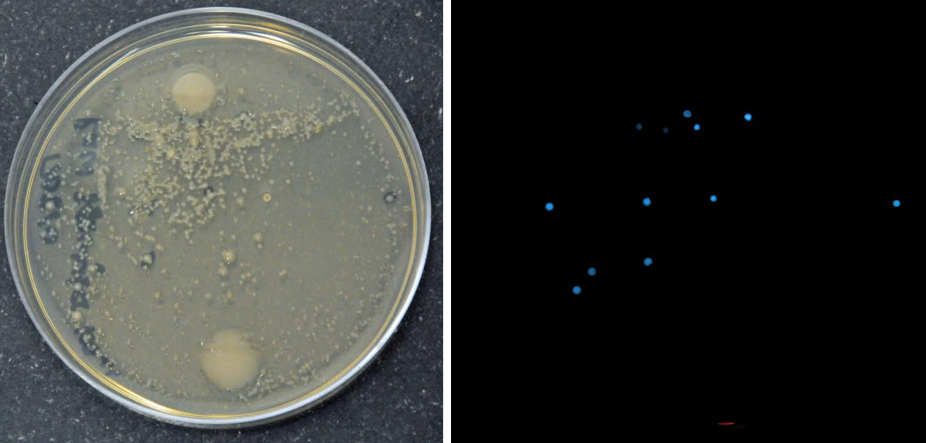
In addition to my research I have been getting to do a lot of education and outreach with some awesome groups! The most recent was the USC C-DEBI high school group who wanted to isolate some bioluminescent bacteria from the cove (see picture below). They were so excited to learn about marine microbes and were very cool to work with! I took a couple of days to go back to the mainland after they left, and now I am looking forward to starting new experiments this upcoming week to continue my search for this novel AAnP group!